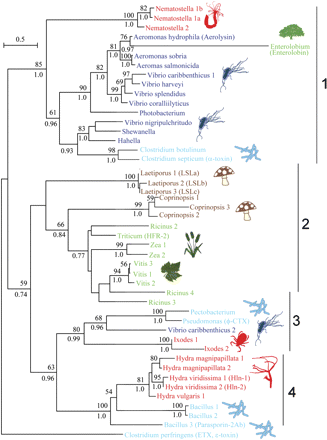
Phylogeny of aerolysin homologues from different kingdoms of life. A ML phylogenetic tree of aerolysin homologues from different organisms was constructed with the WAG model (+F, +G). The organisms that produce the aerolysins are indicated by genus names. In cases where several species of the same genus produce aerolysin-like proteins, the full species names are indicated. The tree is rooted with the epsilon-toxin from Clostridium perfringens. GI numbers of proteins appear in supplementary table 1 (Supplementary Material online.) Bootstrap support values above 50% are indicated above branches and Bayesian posterior probability values above 0.75 appear below branches. Gammaproteobacteria appear in dark blue, Firmicute bacteria in light blue, plants in green, fungi in brown, and animals in red. The corresponding cluster numbers from the clans analysis appear to the left of the tree.




